Epoch 2️⃣4️⃣: This week in ML (+ Bioinformatics 🧬 and Astronomy 🌌)
AlphaFold, Frozen MultiModal Models, ESM, RoseTTA and more ...

AlphaFold leverages novel neural network architectures and training procedures. In summary, the proposed approach to improve accuracy of structure prediction involves: 1) an architecture that jointly embeds multiple sequence alignments (MSA) and pairwise features, 2) a new output representation and loss that improves end-to-end structure prediction, an equivariant attention architecture, a mechanism to iteratively refine predictions using intermediate losses, masked MSA loss to jointly train with structure, self-distillation to learn from unlabelled protein sequences, and self-estimates of accuracy. Overall, AlphaFold incorporates physical and biological knowledge about protein structure and achieves competitive results in the challenging CASP14.
When trained at sufficient scale, auto-regressive language models exhibit the notable ability to learn a new language task after being prompted with just a few examples. Here, the authors present a simple, yet effective, approach for transferring this few-shot learning ability to a multimodal setting (vision and language). Using aligned image and caption data, they train a vision encoder to represent each image as a sequence of continuous embeddings, such that a pre-trained, frozen language model prompted with this prefix generates the appropriate caption. The resulting system is a multimodal few-shot learner, with the surprising ability to learn a variety of new tasks when conditioned on examples, represented as a sequence of multiple interleaved image and text embeddings.
ML in Bioinformatics 🧬
Language models enable zero-shot prediction of the effects of mutations on protein function
Modeling the effect of sequence variation on function is a fundamental problem for understanding and designing proteins. Since evolution encodes information about function into patterns in protein sequences, unsupervised models of variant effects can be learned from sequence data. The approach to date has been to fit a model to a family of related sequences. The conventional setting is limited, since a new model must be trained for each prediction task. The authors show that using only zero-shot inference, without any supervision from experimental data or additional training, protein language models capture the functional effects of sequence variation, performing at state-of-the-art. Link to the code.
Accurate prediction of protein structures and interactions using a three-track neural network
DeepMind presented remarkably accurate predictions at the recent CASP14 protein structure prediction assessment conference. RosettaCommons explored network architectures incorporating related ideas and obtained the best performance with a three-track network in which information at the 1D sequence level, the 2D distance map level, and the 3D coordinate level is successively transformed and integrated. The three-track network produced structure predictions with accuracies approaching those of DeepMind in CASP14, enabling the rapid solution of challenging X-ray crystallography and cryo-EM structure modeling problems, and provides insights into the functions of proteins of currently unknown structure. Link to the code.
Astroinformatics 🌌
The point spread function (PSF) reflects states of a telescope and plays an important role in development of data processing methods, such as PSF based astrometry, photometry and image restoration. However, for wide field small aperture telescopes (WFSATs), estimating PSF in any position of the whole field of view is hard, because aberrations induced by the optical system are quite complex and the signal to noise ratio of star images is often too low for PSF estimation. In this paper, the authors further develop their deep neural network (DNN) based PSF modelling method and show its applications in PSF estimation. During the telescope alignment and testing stage, their method collects system calibration data through modification of optical elements within engineering tolerances (tilting and decentering). Then they use these data to train a DNN (TelNet). After training, the TelNet can estimate PSF in any field of view from several discretely sampled star images. They use both simulated and experimental data to test performance of their method.
Reconstruction of the Density Power Spectrum from Quasar Spectra using Machine Learning
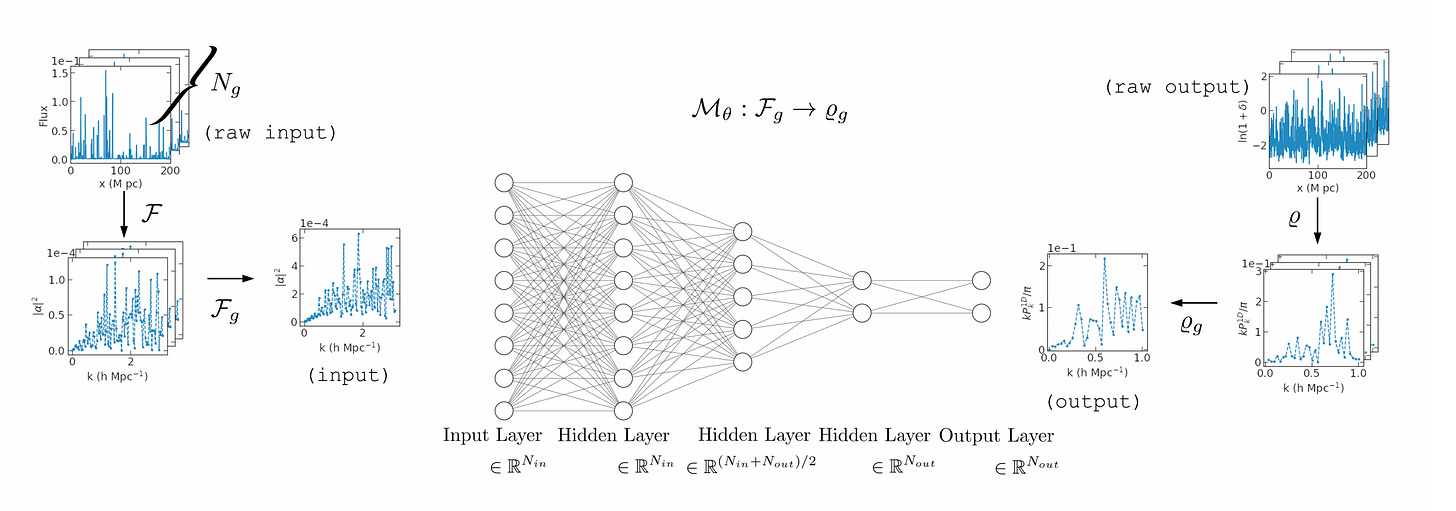
The authors describe a novel end-to-end approach using Machine Learning to reconstruct the power spectrum of cosmological density perturbations at high redshift from observed quasar spectra. State-of-the-art cosmological simulations of structure formation are used to generate a large synthetic dataset of line-of-sight absorption spectra paired with 1-dimensional fluid quantities along the same line-of-sight, such as the total density of matter and the density of neutral atomic hydrogen. With this dataset, they build a series of data-driven models to predict the power spectrum of total matter density. This work provides a foundation for developing methods to analyse very large upcoming datasets with the next-generation observational facilities.
Explainable, Interpretable, Bias and Ethics in AI
Interesting Events and News 📰
Freelancing in Machine Learning | August 13, 2021 | DataTalks.Club
Articles and Resources 📃 I liked
Recommended Podcasts 🎧
Gradient Dissent | Enterprise-scale machine translation with Spence Green, CEO of Lilt
The Real Python Podcast | Planning a Faster Future at the Python Language Summit
TWIML AI Podcast | ML Innovation in Healthcare with Suchi Saria - #501
Towards Data Science | 92. Daniel Filan - Peering into neural nets for AI safety
Lex Fridman Podcast | #201 – Konstantin Batygin: Planet 9 and the Edge of Our Solar System