Epoch 1️⃣4️⃣: This week in ML (+ Bioinformatics 🧬 and Astronomy 🌌)
GANcraft, ROaD, Enformer, generative models for protein design
GANcraft, an unsupervised neural rendering framework for generating photorealistic images of large 3D block worlds such as those created in Minecraft. The method takes a semantic block world as input, where each block is assigned a label such as dirt, grass, tree, sand, or water. They represent the world as a continuous volumetric function and train our model to render view-consistent photorealistic images from arbitrary viewpoints, in the absence of paired ground truth real images for the block world. In addition to camera pose, GANcraft allows user control over both scene semantics and style.
Abstract: In this paper, the explore multi-task learning (MTL) as a second pre-training step to learn enhanced universal language representation for transformer language models. They use the MTL enhanced representation across several natural language understanding tasks to improve performance and generalization. Moreover, they incorporate knowledge distillation (KD) in MTL to further boost performance and devise a KD variant that learns effectively from multiple teachers. By combining MTL and KD, they propose Robustly Optimized and Distilled (ROaD) modeling framework. We use ROaD together with the ELECTRA model to obtain state-of-the-art results for machine reading comprehension and natural language inference.
ML + Bioinformatics 🧬
Effective gene expression prediction from sequence by integrating long-range interactions
Abstract: The next phase of genome biology research requires understanding how DNA sequence encodes phenotypes, from the molecular to organismal levels. How noncoding DNA determines gene expression in different cell types is a major unsolved problem, and critical downstream applications in human genetics depend on improved solutions. Here, the authors report substantially improved gene expression prediction accuracy from DNA sequence through the use of a new deep learning architecture called Enformer that is able to integrate long-range interactions (up to 100 kb away) in the genome. This improvement yielded more accurate variant effect predictions on gene expression for both natural genetic variants and saturation mutagenesis measured by massively parallel reporter assays. Notably, Enformer outperformed the best team on the critical assessment of genome interpretation (CAGI5) challenge for noncoding variant interpretation with no additional training.
Protein sequence design with deep generative models
Abstract: Protein engineering seeks to identify protein sequences with optimized properties. When guided by machine learning, protein sequence generation methods can draw on prior knowledge and experimental efforts to improve this process. In this review, the authors highlight recent applications of machine learning to generate protein sequences, focusing on the emerging field of deep generative methods.
Astroinformatics 🌌
Reducing the complexity of chemical networks via interpretable autoencoders
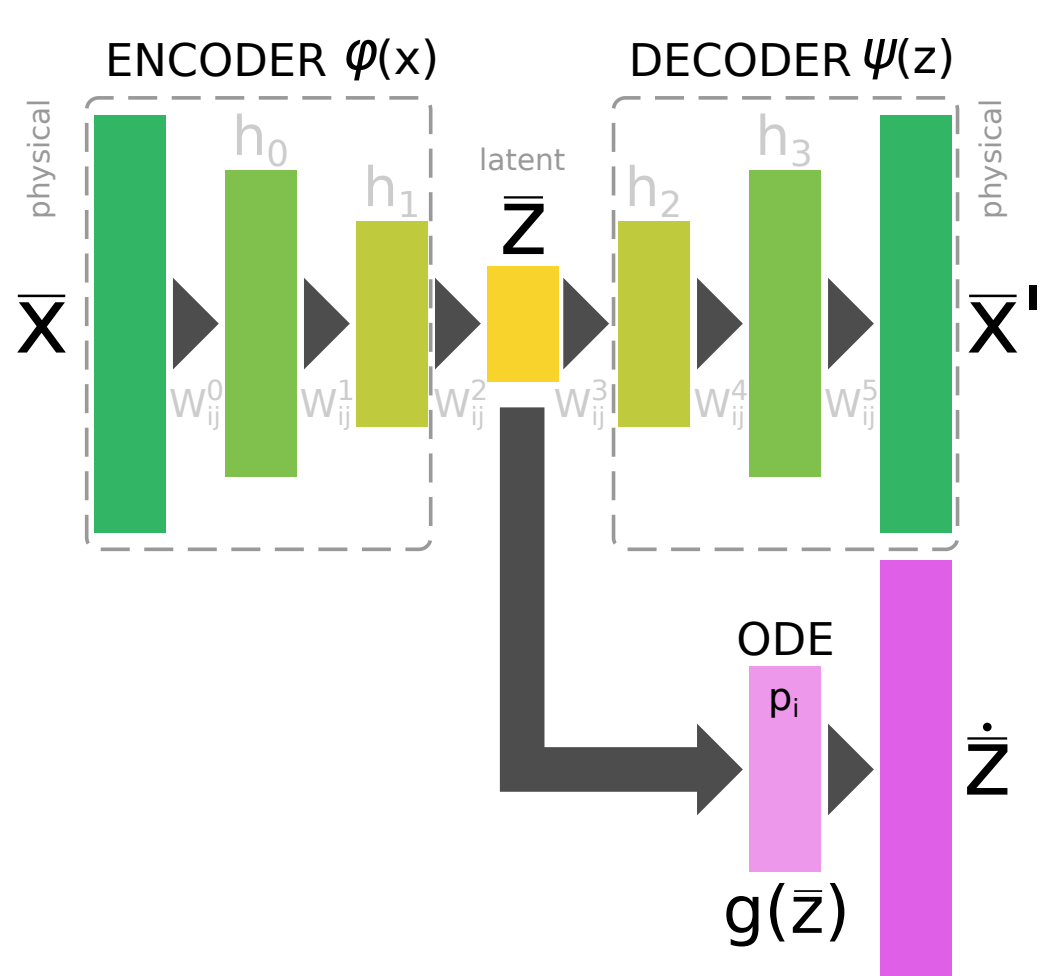
Abstract: In many astrophysical applications, the cost of solving a chemical network represented by a system of ordinary differential equations (ODEs) grows significantly with the size of the network, and can often represent a significant computational bottleneck, particularly in coupled chemo-dynamical models. Although standard numerical techniques and complex solutions tailored to thermochemistry can somewhat reduce the cost, more recently, machine learning algorithms have begun to attack this challenge via data-driven dimensional reduction techniques. In this work, the authors present a new class of methods that take advantage of machine learning techniques to reduce complex data sets (autoencoders), the optimization of multi-parameter systems (standard backpropagation), and the robustness of well-established ODE solvers to to explicitly incorporate time-dependence. This new method allows us to find a compressed and simplified version of a large chemical network in a semi-automated fashion that can be solved with a standard ODE solver, while also enabling interpretability of the compressed, latent network.
A Thesaurus for Common Priors in Gravitational-Wave Astronomy
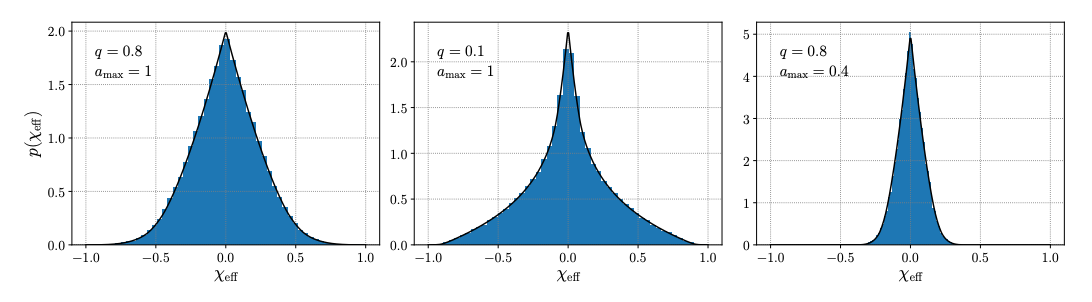
Abstract: In gravitational-wave data analysis, we regularly work with a host of non-trivial prior probabilities on compact binary masses, redshifts, and spins. We must regularly manipulate these priors, computing the implied priors on a transformed basis of parameters or reweighing posterior samples from one prior to another. Here, the authors details some common manipulations, presenting a table of Jacobians with which to transform priors between mass parametrizations, describing the conversion between source- and detector-frame priors, and deriving analytic expressions for priors on the "effective spin" parameters regularly invoked in gravitational-wave astronomy.
Explainable, Interpretable, Bias and Ethics in AI
Interesting Events and News 📰
UnderstandingDL | April 19 - July 2 2021
Articles and Resources 📃 I liked
How I publish articles to all developer platforms (and my private blog) in one shot
Branch Specialization ⭐️ ⭐️
🎥 A survey on generative adversarial networks: fundamentals and recent advances
EPSRC Centre for Doctoral Training in Robotics and Autonomous Systems Newsletter
Characterizing signal propagation to close the performance gap in unnormalized ResNets
Recommended Podcasts 🎧
Gradient Dissent | Nanit's Nimrod Shabtay on deployment and monitoring
The AI Health Podcast | COVID-19 and Racial Inequality with Microsoft Research's Dr. Emma Pierson
The Robot Brains Podcast | Cade Metz talks about how AI took over the world